Fleur Spin-Spiral Dispersion Converge workchain
Current version: 0.2.0
Class:
FleurSSDispConvWorkChainString to pass to the
WorkflowFactory():fleur.ssdisp_convWorkflow type: Scientific workchain, self-consistent subgroup
Aim: Calculate spin-spiral energy dispersion over given q-points converging all the q_points.
Contents
Import Example:
from aiida_fleur.workflows.ssdisp_conv import FleurSSDispConvWorkChain
#or
WorkflowFactory('fleur.ssdisp_conv')
Description/Purpose
This workchain calculates spin spiral energy dispersion over a given set of q-points. Resulting energies do not contain terms, corresponding to DMI energies. To take into account DMI, see the Fleur Dzyaloshinskii–Moriya Interaction energy workchain documentation.
In this workchain the force-theorem is employed which means the workchain converges
a reference charge density first
and then submits a single FleurCalculation with a <forceTheorem> tag. However, it is possible
to specify inputs to use external pre-converged charge density to use it as a reference.
The task of the workchain us to calculate the energy difference between two or several structures having a different magnetisation profile:
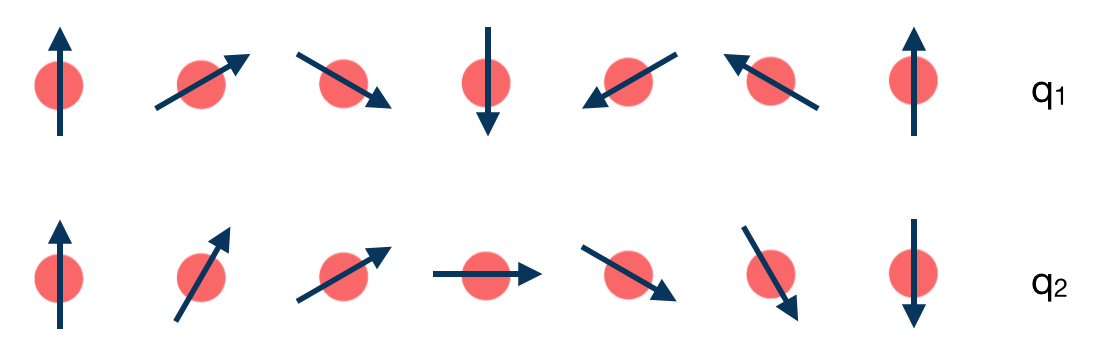
To do this, the workchain employs the force theorem approach:
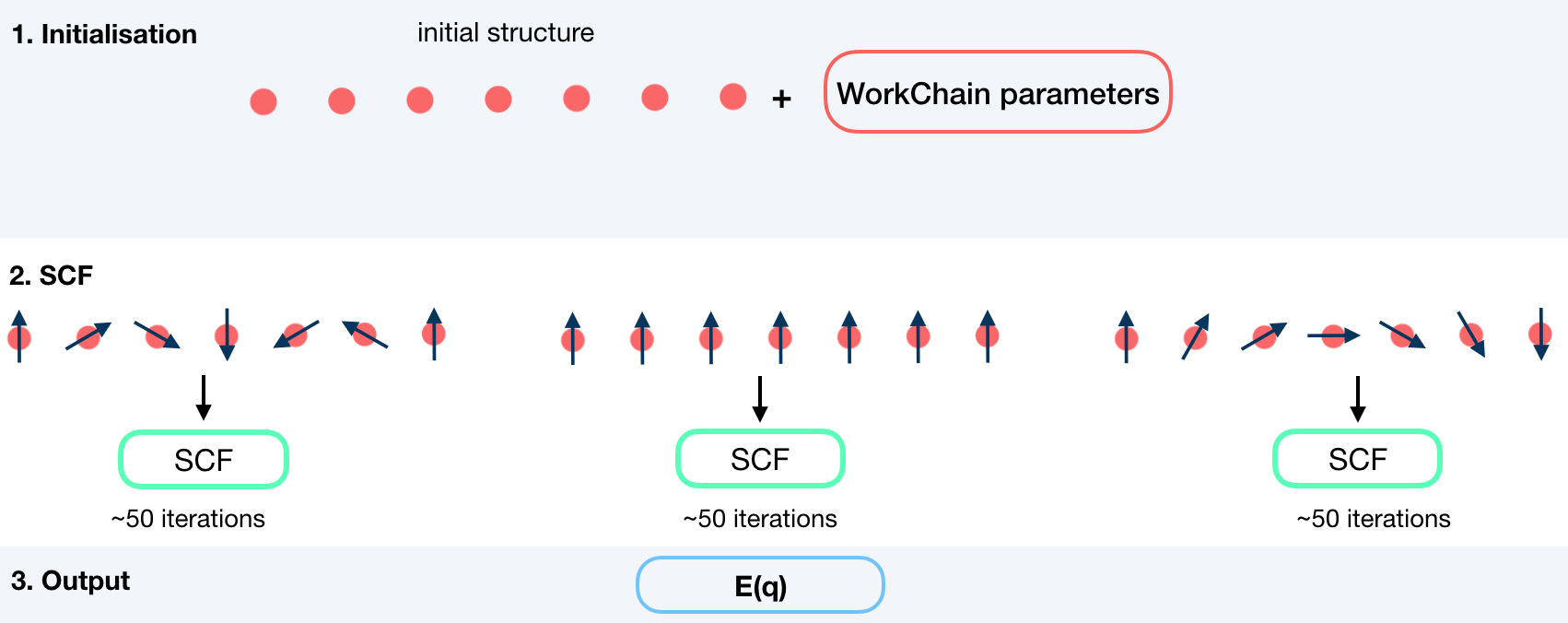
Input nodes
The FleurSSDispWorkChain employs
exposed feature of the AiiDA, thus inputs for the nested
SCF workchain should be passed in the namespace
scf.
name |
type |
description |
required |
|---|---|---|---|
scf |
namespace |
inputs for nested SCF WorkChain |
yes |
wf_parameters |
Settings of the workchain |
no |
Workchain parameters and its defaults
wf_parameters
wf_parameters: Dict - Settings of the workflow behavior. All possible
keys and their defaults are listed below:
# -*- coding: utf-8 -*-
'beta': {'all': 1.57079}, # see the description below
'q_vectors': {'label': [0.0, 0.0, 0.0], # sets q_points to calculate
'label2': [0.125, 0.0, 0.0]
}
'suppress_symmetries': False # True if use no symmetries
beta is a python dictionary containing a key: value pairs. Each pair sets beta parameter
in an inp.xml file. key specifies the atom label to change, key equal to ‘all’ sets all
atoms groups. For example,
'beta' : {'222' : 1.57079}
changes
<atomGroup species="Fe-1">
<filmPos label=" 222">.0000000000 .0000000000 -11.4075100502</filmPos>
<force calculate="T" relaxXYZ="TTT"/>
<nocoParams l_relax="F" alpha=".00000000" beta="0.00000" b_cons_x=".00000000" b_cons_y=".00000000"/>
</atomGroup>
to:
<atomGroup species="Fe-1">
<filmPos label=" 222">.0000000000 .0000000000 -11.4075100502</filmPos>
<force calculate="T" relaxXYZ="TTT"/>
<nocoParams l_relax="F" alpha=".00000000" beta="1.57079" b_cons_x=".00000000" b_cons_y=".00000000"/>
</atomGroup>
Note
beta actually sets a beta parameter for a whole atomGroup. It can be that the atomGroup correspond to several atoms and beta switches sets beta for atoms that was not intended to change. You must be careful and make sure that several atoms do not correspond to a given specie.
q_vectors is a python dictionary (key: value pairs). The key can be any string which
sets a label of the q-vector. value must be a list of 3 values: $$q_x, q_y, q_z$$.
Output nodes
out:Dict- Information of workflow results like success, last result node, list with convergence behavior{ "energies": { "label": 0.0, "label2": 0.014235119451769 }, "energy_units": "eV", "errors": [], "failed_labels": [], "info": [], "q_vectors": { "label": [ 0.0, 0.0, 0.0 ], "label2": [ 0.125, 0.0, 0.0 ] }, "warnings": [], "workflow_name": "FleurSSDispConvWorkChain", "workflow_version": "0.1.0" }Resulting Spin Spiral energies are listed according to given labels.
Layout
SSDisp converge always starts with a structure and a list of q-vectors to calculate. There is no way to continue from pre-converged charge density.
Error handling
A list of implemented exit codes:
Code |
Meaning |
|---|---|
230 |
Invalid workchain parameters |
340 |
Convergence SSDisp calculation failed for all q-vectors |
341 |
Convergence SSDisp calculation failed for some q-vectors |
Example usage
# -*- coding: utf-8 -*- from aiida.orm import load_node, Dict from aiida.engine import submit from aiida_fleur.workflows.ssdisp_conv import FleurSSDispConvWorkChain fleur_code = load_node(FLEUR_PK) inpgen_code = load_node(INPGEN_PK) structure = load_node(STRUCTURE_PK) wf_para = Dict(dict={'beta': {'all': 1.57079}, 'q_vectors': {'label': [0.0, 0.0, 0.0], 'label2': [0.125, 0.0, 0.0] } }) options = Dict(dict={'resources': {'num_machines': 1, 'num_mpiprocs_per_machine': 24}, 'queue_name': 'devel', 'custom_scheduler_commands': '', 'max_wallclock_seconds': 60*60}) parameters = Dict(dict={'atom': {'element': 'Pt', 'lmax': 8 }, 'atom2': {'element': 'Fe', 'lmax': 8, }, 'comp': {'kmax': 3.8, }, 'kpt': {'div1': 20, 'div2': 24, 'div3': 1 }}) wf_para_scf = {'fleur_runmax': 2, 'itmax_per_run': 120, 'density_converged': 0.2, 'mode': 'density' } wf_para_scf = Dict(dict=wf_para_scf) options_scf = Dict(dict={'resources': {'num_machines': 2, 'num_mpiprocs_per_machine': 24}, 'queue_name': 'devel', 'custom_scheduler_commands': '', 'max_wallclock_seconds': 60*60}) inputs = {'scf': {'wf_parameters': wf_para_scf, 'structure': structure, 'calc_parameters': parameters, 'options': options_scf, 'inpgen': inpgen_code, 'fleur': fleur_code }, 'wf_parameters': wf_para, } res = submit(FleurSSDispConvWorkChain, **inputs)